Research


Volatile chemicals in the environment (odorants) are detected by olfactory sensory neurons in the nasal cavity. These sensory neurons send signals directly to the olfactory bulb, the first region of the brain that processes olfactory information. Each sensory neuron expresses only 1 out of >1,000 odorant receptor genes, and neurons that express the same receptor project their axons to specific structures in the bulb called glomeruli, setting up a spatial representation of chemical information in the brain.
Because of this organization, the olfactory system is arguably the most genetically tractable of the senses. The function and wiring of different neuronal populations are defined by the expression of single genes and can thus be manipulated genetically with relative ease. Olfaction also offers a unique view into how the brain represents sensory information, and how individual genes influence neuronal connections and behavior. The lab is currently working on several projects.
Molecular organization of the olfactory map
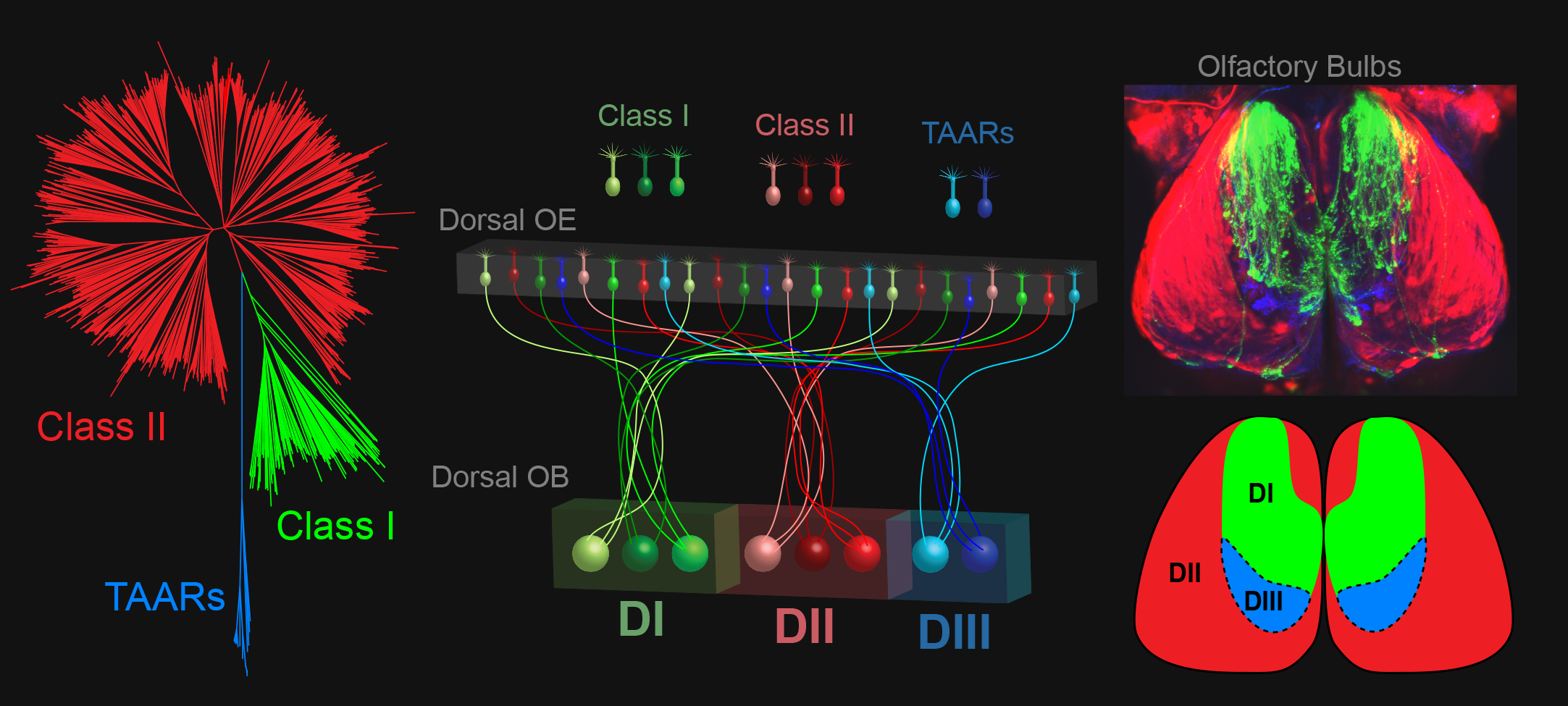
Work from the Bozza lab has shown that the odorant receptor repertoire is broadly mapped on the surface of the olfactory bulb. Mammalian odorant receptors can be divided into three phylogenetically distinct classes (tree above), canonical Class I and Class II odorant receptors (green and red), and a small family of Trace Amine Associated Receptors, or TAARs (blue). We have discovered that all three classes of mammalian olfactory receptors are mapped to discrete domains in the dorsal olfactory bulb. We are currently determining how these projections form and how they function in sensory processing.
Mechanisms of monoallelic gene expression
How does each olfactory sensory neuron in the mouse exclusively express 1 allele of 1 receptor gene out of 2,000 alleles? Studying olfactory gene regulation is daunting because of the large number of genes and because they are scattered across many genomic clusters throughout the genome. To simplify the problem, we are using the Trace Amine-Associated Receptors as a model to understand the genetic basis for receptor gene choice.
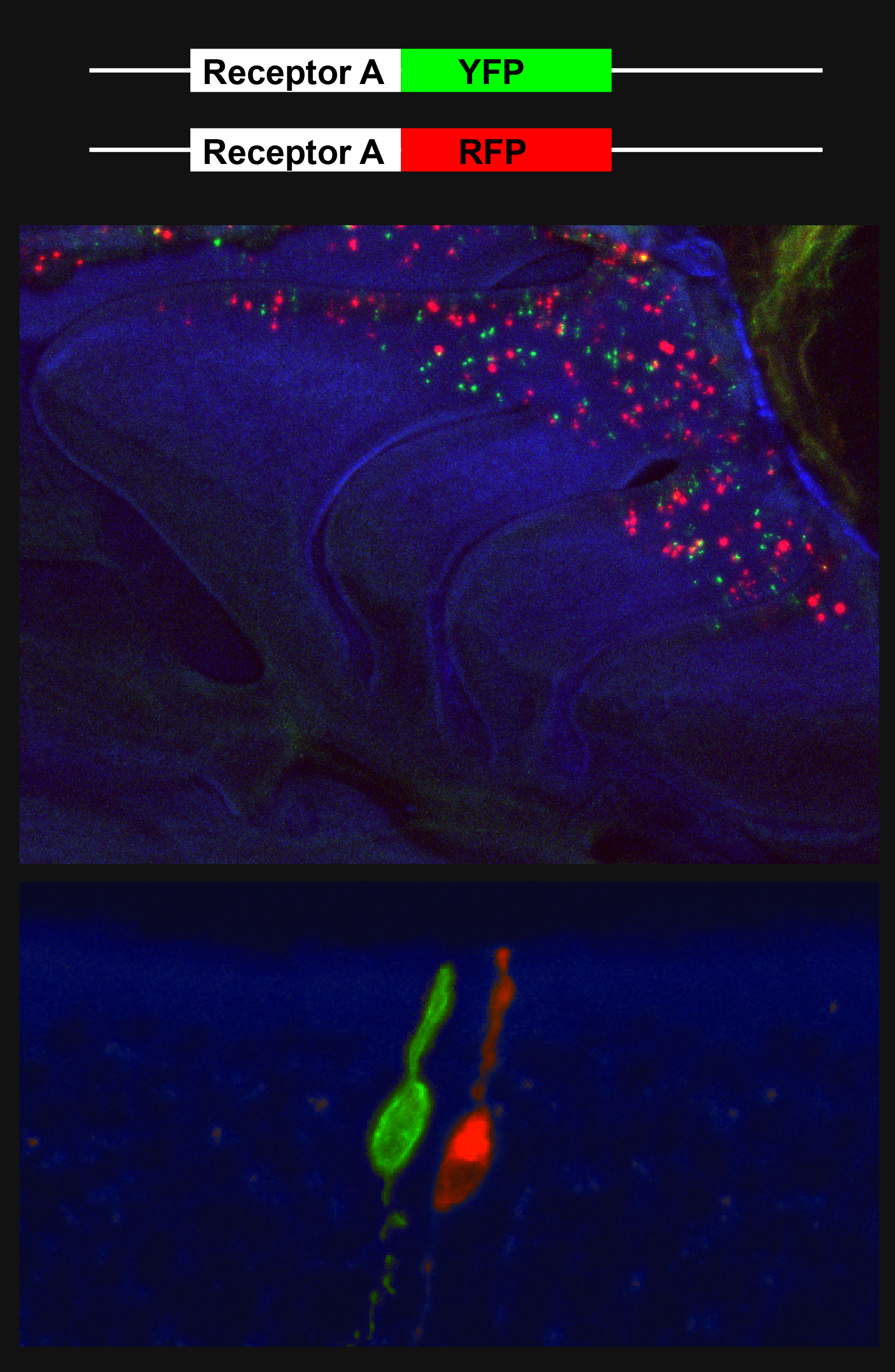
There are only 14 TAAR genes in mouse and they are found in a single, uninterrupted gene cluster. The TAARs share functional similarity with the main classes of canonical odorant receptors and they are monoallelically expressed. The figure shows monoallelic expression of differentially tagged TAAR genes, shown in green and red.
Using CRISPR-based gene targeting, transgenics, transcriptomics and epigenetic analyses in mice, we are studying the function of cis-acting enhancers that control TAAR gene expression.
Genetics of neuronal diversification and axon guidance
The olfactory system is one of very few sites that show adult neurogenesis in mammals. Stem cells in the olfactory epithelium give rise to functionally distinct populations of olfactory sensory neurons, which then grow axons to connect to the olfactory bulb throughout life. Surprinsingly, sensory neurons that express different odorant receptors are transcriptionally unique. The expressed GPCR dominates their transcriptional identity. This identity is thought to influence axon guidance to allow axons to converge to glomeruli in a receptor-specific way.
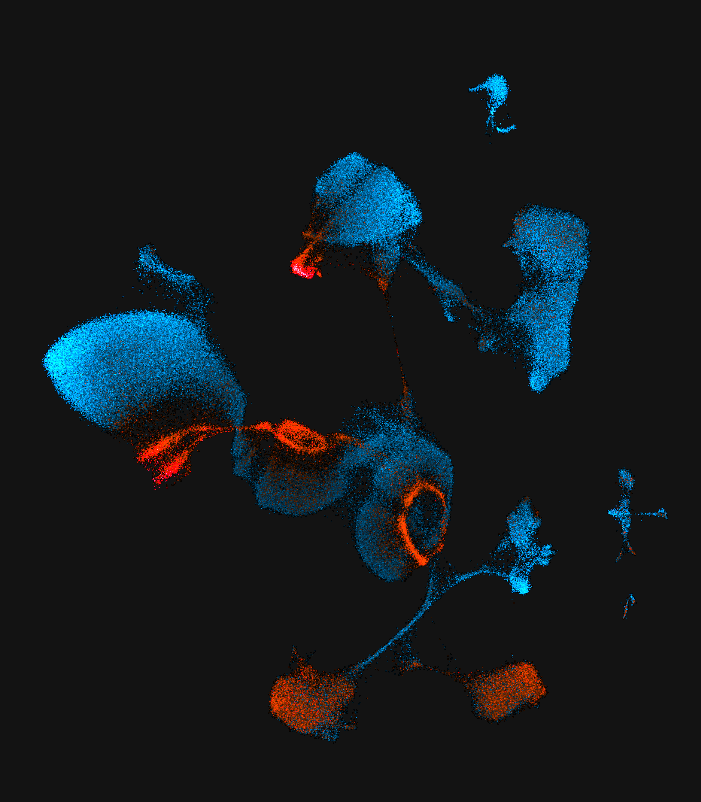
We are using single cell genomics aproaches to understand how transcriptionally diverse sensory neuron cell types arise during neurogenesis and how transcriptional diversity biases receptor gene choice and axon guidance
How olfactory receptor genes contribute to behavior
Mammals retain large families of olfactory receptor genes (typically in the thousands). What determines how many receptors a species has? How are the genes retained during evolution, and how do individual receptors contribute to defined percepts and behaviors?
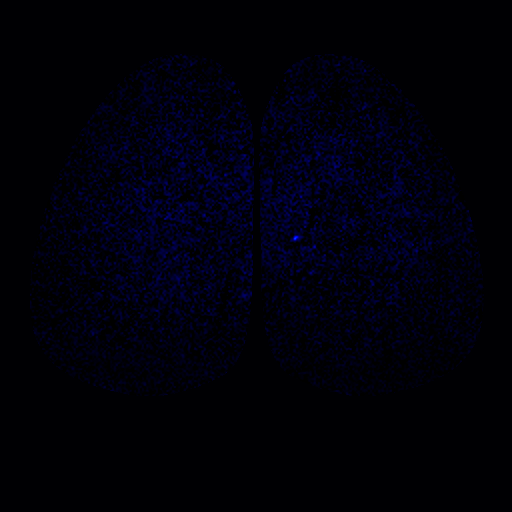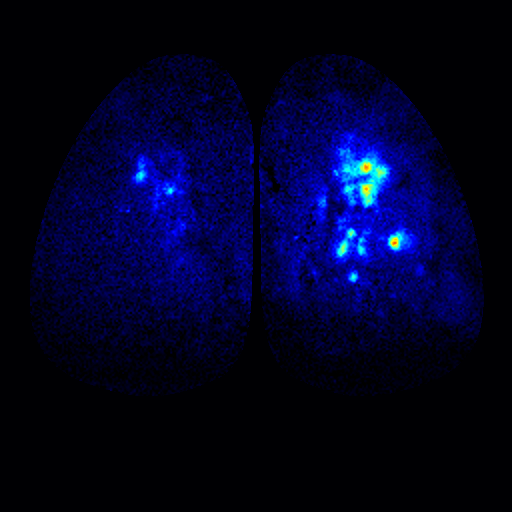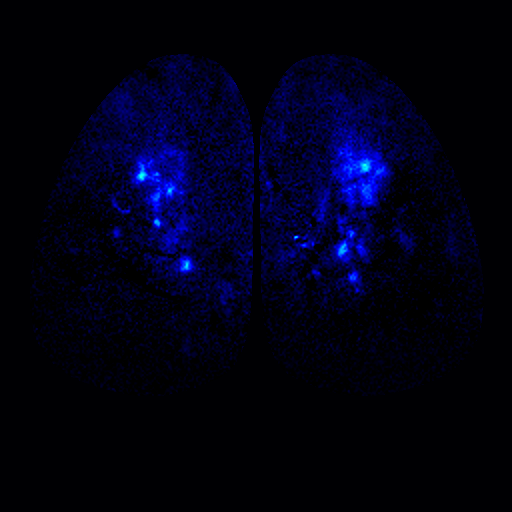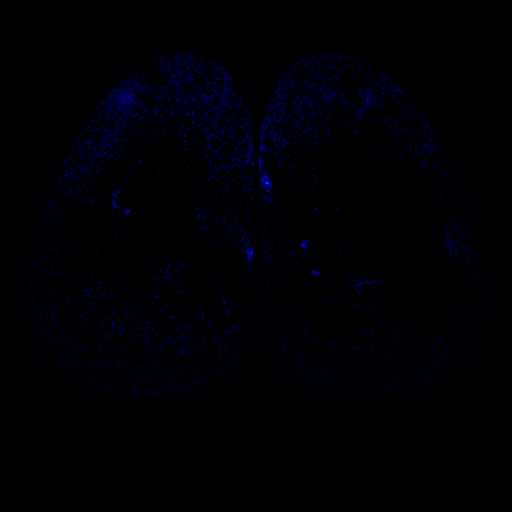
To address these questions, we have been studying the TAARs: a small evolutionarily conserved family of olfactory receptors. We are characterizing the function of TAARs using electrophysiology, imaging, and behavior in gene targeted mice. We have shown that these receptors are highly sensitive to amines and that a majority of the TAARs map to a subset of amine-selective glomeruli in the doral olfactory bulb (below). Using advanced mouse genetics, we have generated mice that lack all TAAR genes, or specific combinations of TAAR genes. Our recent data demonstrate that individual TAARs impact perception in two ways: sensitivity and ethologically relevant behaviors.
Sensitivity: We have shown that the TAARs are the most sensitive receptors for a class of chemicals called amines, and that they set the sensitivity of mice to individual amine odors. Knocking out the TAARs makes mice less sensitive to amines. Identifying high-affinity receptors that impact sensitivity is a landmark finding. This has led us to search for other highly sensitive olfactory receptors in order to study the molecular basis for olfactory sensitivity and with Dima Rinberg's lab at NYU, general principles of odor coding.
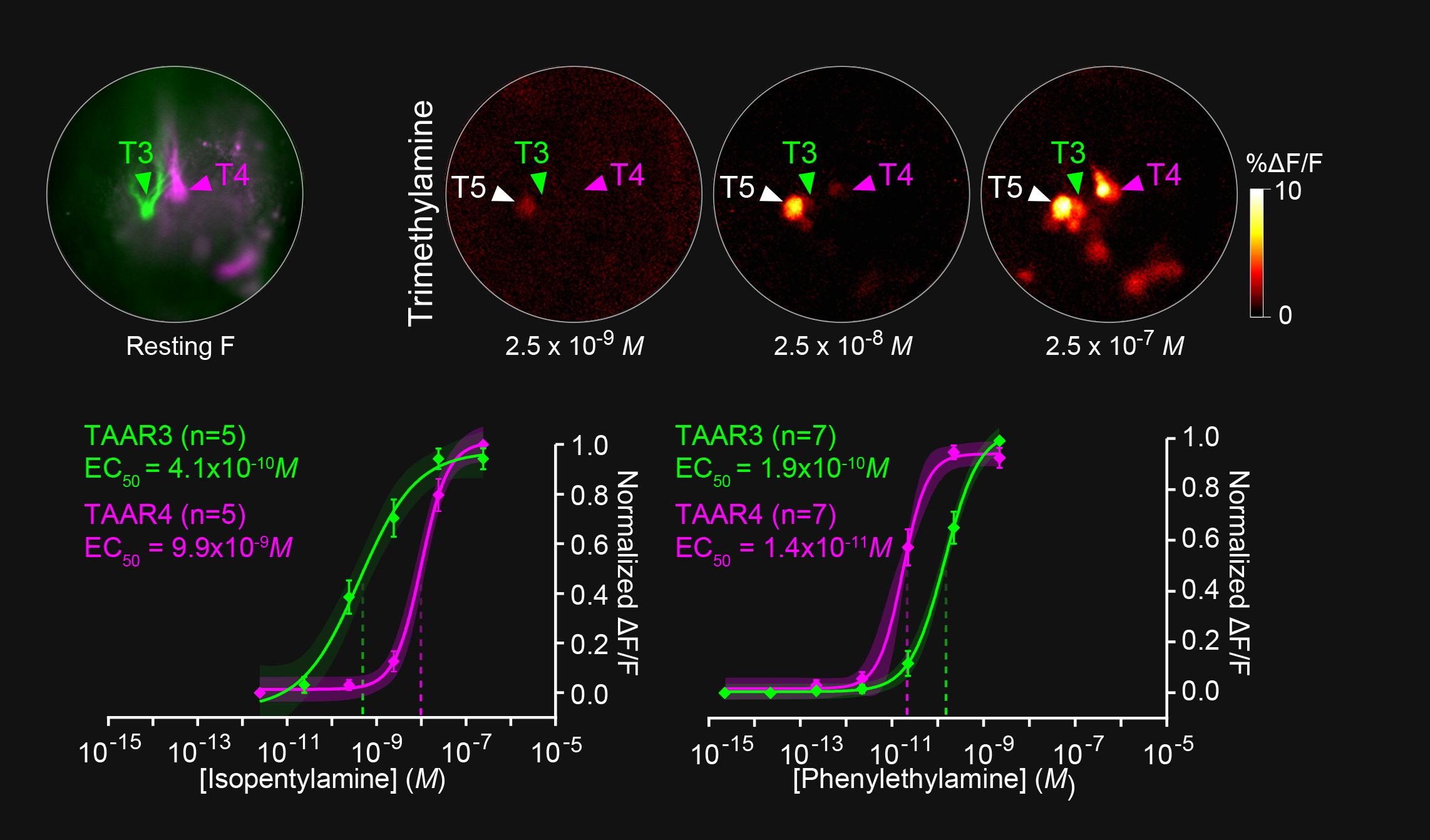
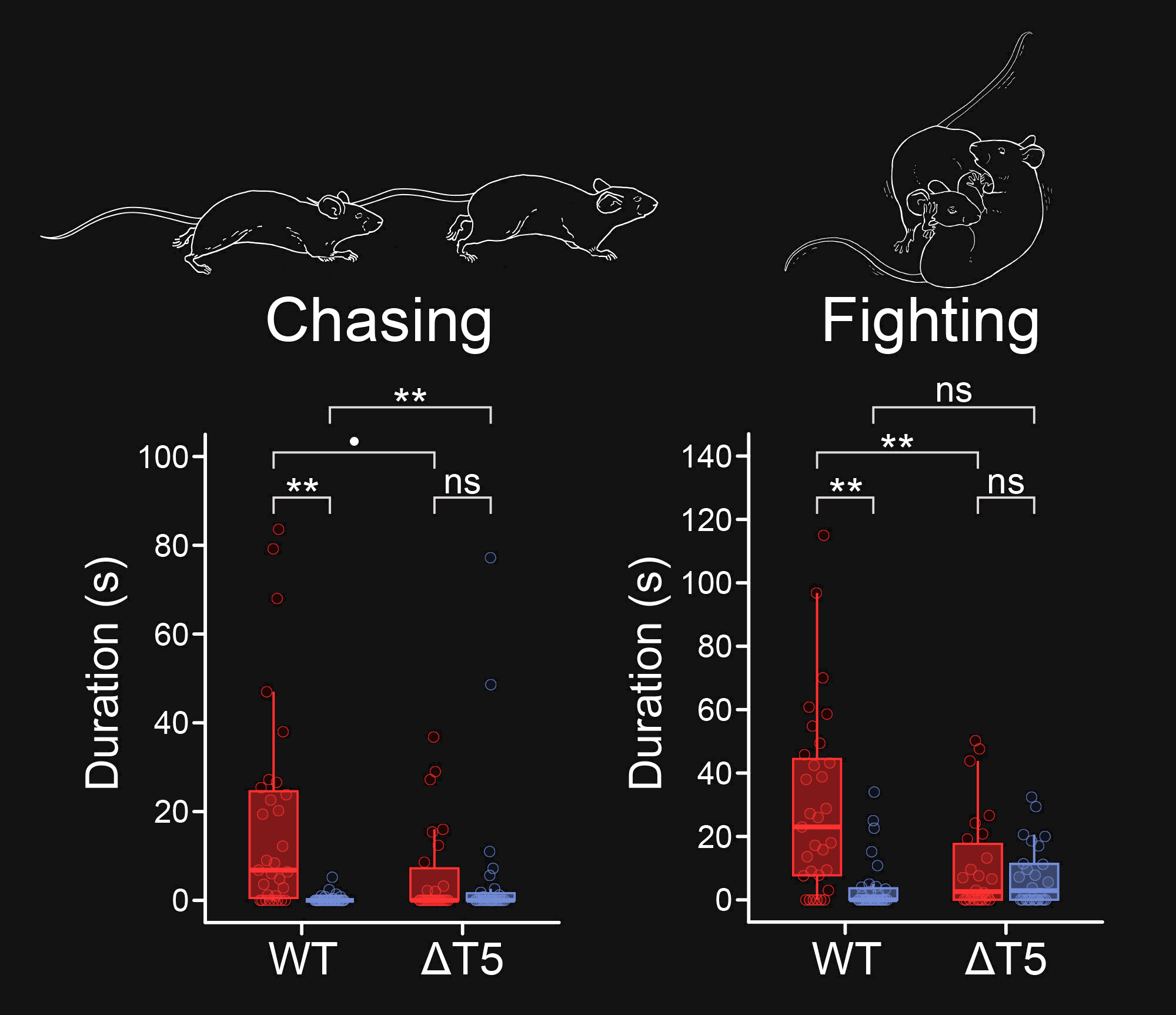
Ethologically Relevant Behaviors: Chemical cues like the amines influence critical behaviors in mammals, such as mating, feeding, maternal behavior, and aggression. We have shown that TAAR4 is critical for innate aversive responses that mice show to the predator odor phenylethylamine. In addition, we have shown that TAAR5 and its ligand trimethylamine (a metabolite produced by the gut microbiomeme) regulate aggression and social dominance in male mice. We are investigating the circuits underlying these behaviors, and whether the TAARs mediate behavioral responses to other ethologically relevant chemosensory stimuli.